🚀 Introduction
proGENTRL is a PyTorch Lightning implementation of Generative Tensorial Reinforcement Learning (GENTRL) — a deep generative model designed to explore chemical space and propose novel molecules with desirable properties, such as synthetic feasibility and biological activity.
This blog post walks through what proGENTRL is, how it works, what’s inside the repository, and how you can get started building and experimenting with it.
Repository: github.com/Bibyutatsu/proGENTRL
🧠 What is GENTRL?
Generative Tensorial Reinforcement Learning (GENTRL) is a two-stage model developed for de novo small-molecule design. The original research demonstrated that GENTRL could:
- Optimize synthetic feasibility, novelty, and biological activity of molecules.
- Generate novel small-molecule inhibitors for the DDR1 kinase target within a matter of weeks.
The model combines a Variational Autoencoder (VAE) with reinforcement learning (RL). Initially, the VAE is trained on a dataset of molecules, learning a smooth latent representation of chemical space. Then, an RL agent explores this latent space to find points that decode into molecules with high reward as defined by property-based scoring.
Research Paper: Deep learning enables rapid identification of potent DDR1 kinase inhibitors - Nature Biotechnology, 2019
🧠 Why It Matters
Chemical space — the total set of possible small molecules — is astronomically large. Traditional in-silico screening methods can only explore a tiny fraction of this. Models like GENTRL use deep learning to generalize learning from existing molecules and guide exploration toward regions that satisfy multiple objectives (like drug-likeness, activity against a target, or synthesis feasibility).
This makes such models powerful tools in drug discovery, materials design, and other chemistry-driven fields.
📦 What’s Inside the proGENTRL Repository
Here’s a high-level look at the repository structure:
proGENTRL/
├─ images/ # Images shown in README
├─ progentrl/ # Core model and training implementation
├─ Example.ipynb # Demo notebook with workflow
├─ README.md # Project information & installation
├─ setup.py # Install script
└─ LICENSE # MIT License
Key Components
| Component | Description |
|---|---|
| VAE Module | Encodes/decodes SMILES strings to/from latent space |
| RL Module | Reinforcement learning for latent space optimization |
| Tokenizer | Custom SMILES tokenizer for molecular strings |
| Trainer | PyTorch Lightning training loops |
🛠️ Installation & Setup
Follow these steps to get proGENTRL running:
Step 1 — Create Conda Environment
Install RDKit, which is required for molecule handling:
conda create -c rdkit -n progentrl-env rdkit
conda activate progentrl-env
Step 2 — Install proGENTRL
Option A: Install via pip:
pip install progentrl
Option B: Install from source:
git clone https://github.com/Bibyutatsu/proGENTRL.git
cd proGENTRL
python3 setup.py install
Step 3 — Install PyTorch
Install PyTorch with the appropriate CUDA version:
python3 -m pip uninstall torch torchvision
conda install pytorch torchvision cudatoolkit=11.3 -c pytorch
Replace cudatoolkit=11.3 with your CUDA version.
Step 4 — (Optional) Setup Jupyter Kernel
python3 -m pip install ipykernel
python3 -m ipykernel install --user --name progentrl
This makes it easy to run notebooks with the correct environment.
🧪 Example Workflow
The included Example.ipynb demonstrates the core workflow:
┌─────────────────┐ ┌──────────────────┐ ┌─────────────────────────┐ ┌─────────────────┐
│ Pretrain VAE │ ──▶ │ Freeze Weights │ ──▶ │ Reinforcement Learning │ ──▶ │ Sample SMILES │
└─────────────────┘ └──────────────────┘ └─────────────────────────┘ └─────────────────┘
- Pretrain the VAE: Learn a latent representation of molecules.
- Freeze VAE weights: Lock them except for the prior.
- Run Reinforcement Learning: Search the latent space for high-reward vectors.
- Sample SMILES strings: Decode optimized latent points into candidate molecules.
This notebook is the recommended starting point to understand how the model flows from training to molecule generation.
📘 Core Concepts Explained
Variational Autoencoder (VAE)
A VAE encodes molecules (e.g., SMILES strings) into a continuous latent space and decodes back to molecule representations. The learning objective encourages the latent space to be smooth and meaningful.
SMILES Input → Encoder → Latent Space (z) → Decoder → SMILES Output
↓ ↓ ↓
"CCO" → [0.2, -0.5, 1.3, ...] → "CCO"
Reinforcement Learning (RL) Optimization
Once pretrained, the VAE’s weights are mostly frozen. Reinforcement learning is applied on top of the latent representation:
- A reward function guides the agent toward latent points that decode into molecules with desired properties.
- The exploration is driven by a reward score (e.g., drug-likeness, synthetic accessibility).
🧭 Real-World Performance
The original GENTRL research found potent DDR1 inhibitors in a short timeframe — generating molecules with promising activity and validating them experimentally in biochemical and cellular assays.
Generated Molecules Examples
Below are examples of molecules generated by the model:
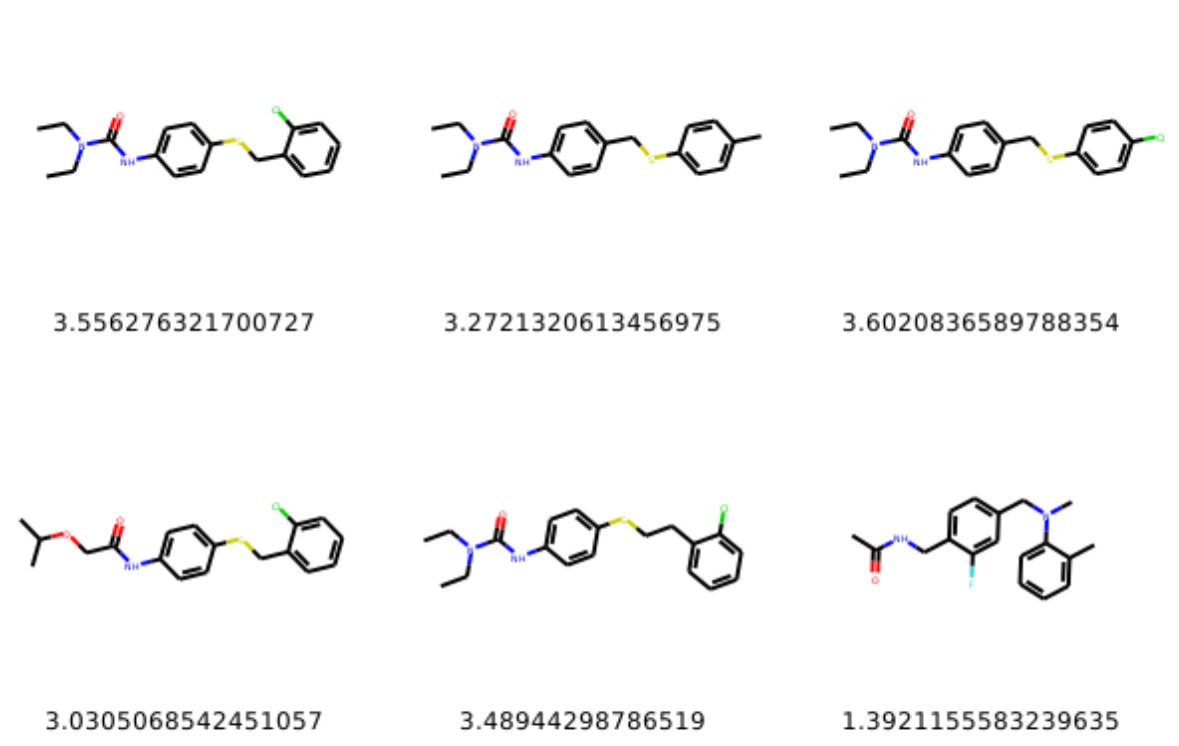
More samples available here
This provides an exciting example of how generative models can accelerate the early stages of drug discovery.
🧠 Extending proGENTRL
Here are ways to enhance the project:
🎯 Better Reward Functions
Design multi-objective reward functions that incorporate:
- Predicted biological target affinity
- Synthetic accessibility scores
- Drug-like metrics (e.g., QED, logP, Lipinski scores)
⚙️ Advanced Generative Models
Beyond VAEs, you can explore:
- Transformer-based generative models — Attention mechanisms for sequential SMILES
- Graph neural networks — Molecular graph representations
- Diffusion models — State-of-the-art generative approach
These can bring richer representations and better optimization.
💡 Final Thoughts
The proGENTRL repository provides a strong starting point to experiment with deep generative models and reinforcement learning in chemistry. Whether you’re exploring de novo drug design or generative modeling, this PyTorch Lightning implementation gives you a practical and modifiable codebase to build from.
Why PyTorch Lightning?
PyTorch Lightning offers:
- Clean separation of research code from engineering
- Multi-GPU support out of the box
- Reproducibility through standardized training loops
- Reduced boilerplate — focus on the model, not infrastructure
📚 Resources
- proGENTRL Repository: github.com/Bibyutatsu/proGENTRL
- Original GENTRL: github.com/insilicomedicine/gentrl
- My GENTRL Fork: github.com/Bibyutatsu/GENTRL
- Research Paper: Nature Biotechnology Publication
| *Originally published: June 7, 2020 | Updated: January 2025* |
